 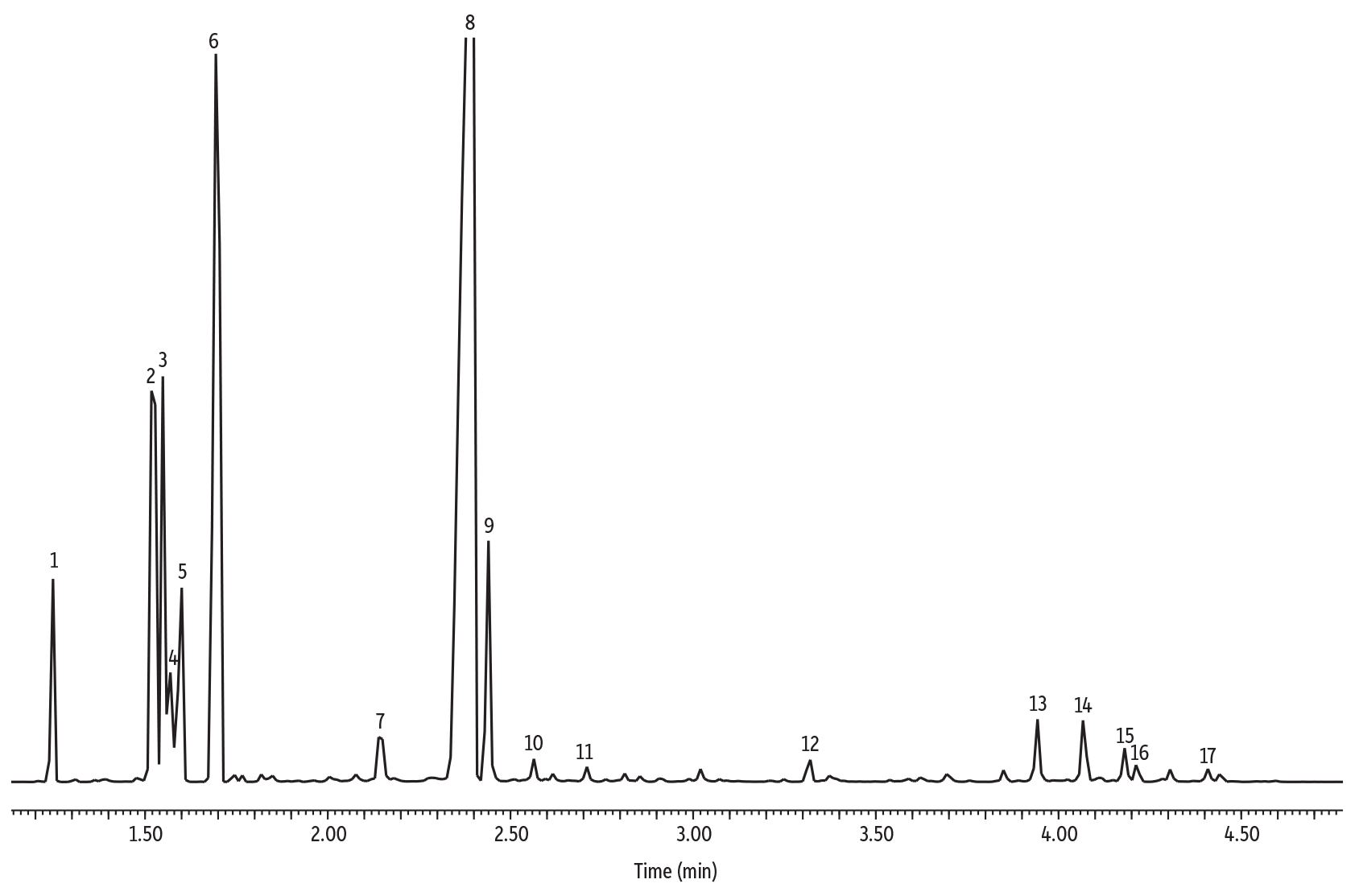
The general procedure for PG isolation and purification has remained relatively unchanged since it was described in the 1960s 9. In Enterococci, substitution of the terminal ᴅ-Ala of the peptide sidechain with ᴅ-Lac confers a greater resistance to the antimicrobial, vancomycin 7, 8. As examples, a common modification is the addition of a C6 acetyl group on the MurNAc that confers resistance by limiting access to the glycan β-1,4 linkages to host-produced lysozyme enzymes which degrade PG 4, 5, 6. 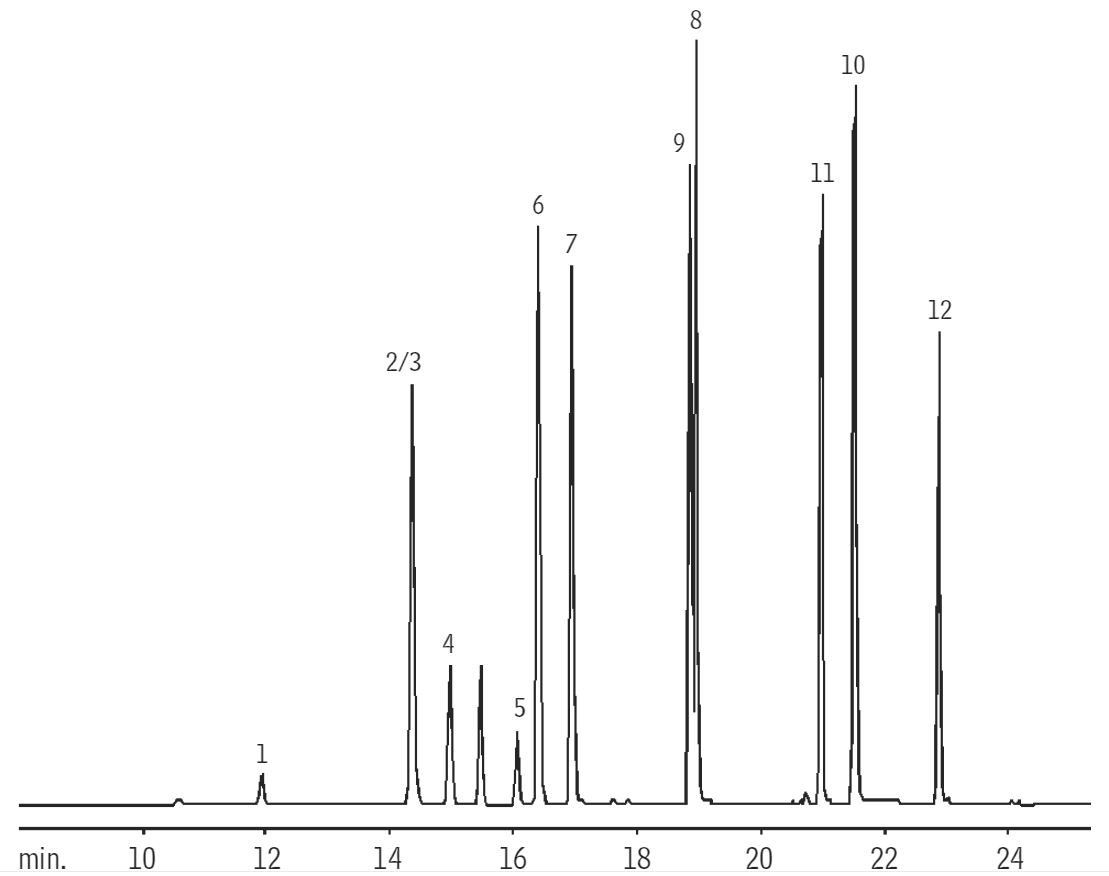
In addition, various modifications of the PG have been shown to confer adaptations specific to the strain, growth conditions, and environmental stress, which have been implicated in cell signalling, antimicrobial resistance, and host immune evasion 4. However, once assembled, there are numerous enzymes produced by the bacteria that modify and/or degrade the PG to carry out a number of cellular processes, including growth and division. Precursors are subsequently incorporated into the mature PG by transglycosylase and transpeptidase enzymes, which produce the glycosidic and peptide bonds, respectively 3. During PG synthesis, precursors are generated in the cytoplasm, and transported across the cytoplasmic membrane by flippases. This crosslinking produces a mesh-like structure that encompasses the entire cell and is often referred to as a sacculus ( Figure 1C). Each MurNAc possesses a short peptide bound at the ᴅ-lactyl residue that can be crosslinked to adjacent disaccharide-linked peptides ( Figure 1 A,B). 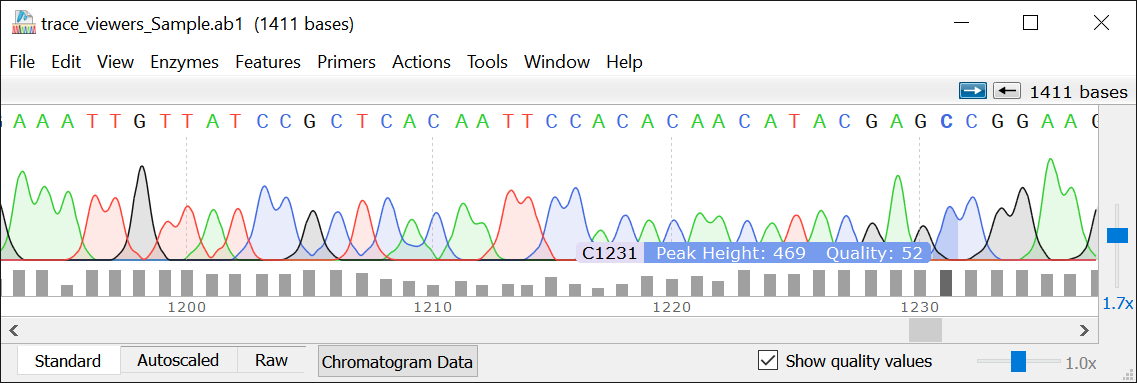
The backbone of PG is composed of alternating β-1,4-linked N-acetyl muramic acid (MurNAc) and N-acetyl glucosamine (GlcNAc) 1, 2. Peptidoglycan (PG) is a defining characteristic of bacteria that serves to maintain cell morphology, while providing structural support for proteins and other cellular components 1, 2. The following protocol describes the purification of peptidoglycan from bacterial cultures, the acquisition of muropeptide intensity data through a liquid chromatograph-mass spectrometer, and the differential analysis of peptidoglycan composition using bioinformatics. 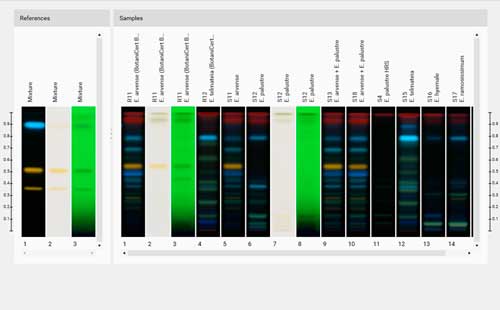
Peptidoglycomics uses advanced mass spectrometry combined with high-powered bioinformatic data analysis to examine peptidoglycan composition in fine detail. To understand the variation within peptidoglycan, the overall structure must be broken down into its constitutive parts (known as muropeptides) and assessed for overall cellular composition. These variations produce a highly dynamic structure that is known to participate in many cellular functions, including growth/division, antibiotic resistance, and host defense avoidance. In addition, there are numerous known variations, modifications, or adaptations to the peptidoglycan that can occur within a bacterial species in response to growth phase and/or environmental stimuli. Although aspects of peptidoglycan structure are fairly conserved across all bacteria, there is also considerable variation between Gram-positives/negatives and between species. Peptidoglycan is an important component of bacterial cell walls and a common cellular target for antimicrobials. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
